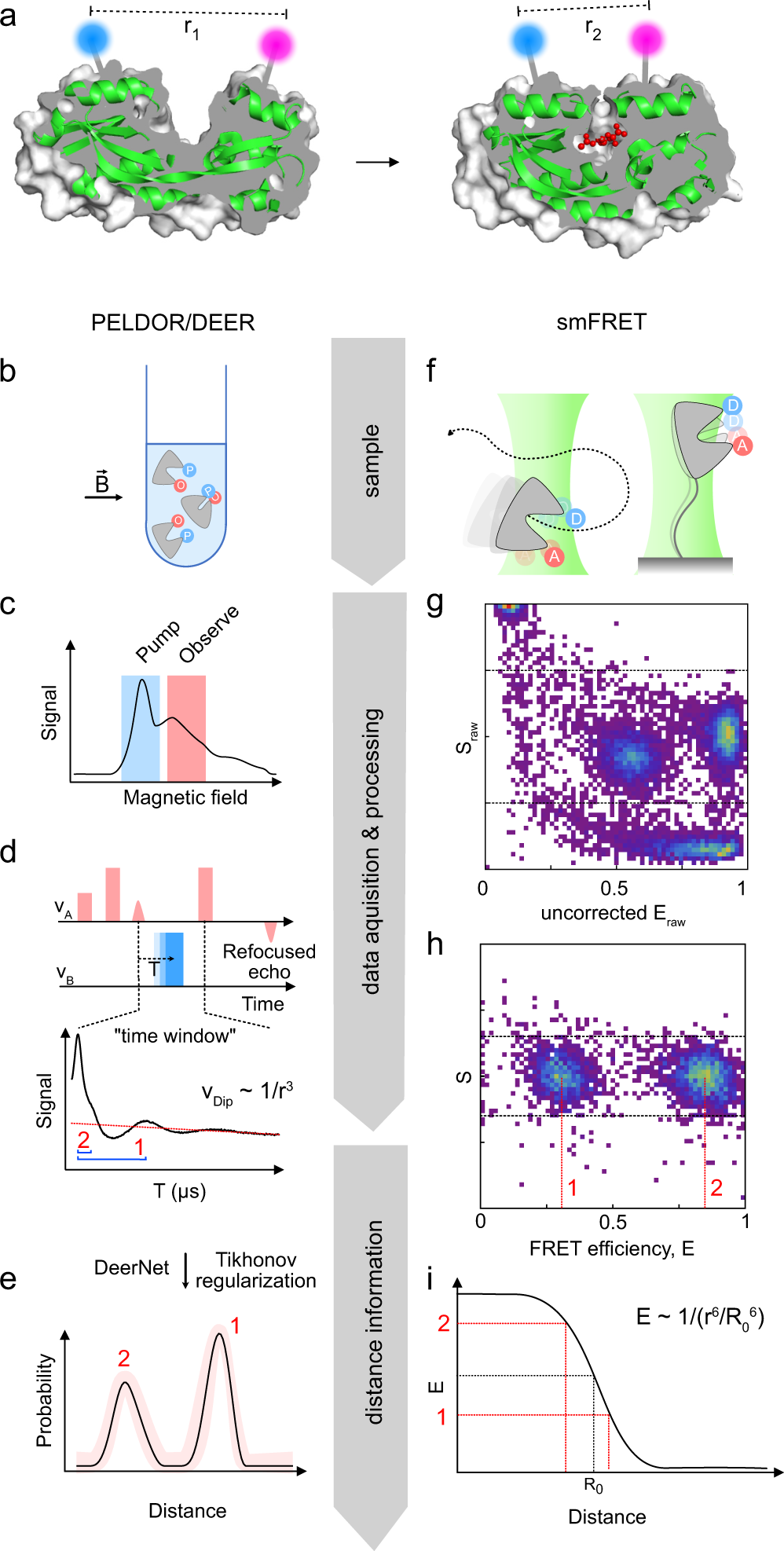
Cross-validation of distance measurements in proteins by PELDOR/DEER and single-molecule FRET | Nature Communications
![SOLVED: For an enzyme following Michaelis-Menten kinetics, the following Vo vs. S data were obtained: [S] 1 micromolar 2 micromolar 4 micromolar 8 micromolar Vo 10 micomolar/min 20 micromolar/min 40 micromlar/min 70 SOLVED: For an enzyme following Michaelis-Menten kinetics, the following Vo vs. S data were obtained: [S] 1 micromolar 2 micromolar 4 micromolar 8 micromolar Vo 10 micomolar/min 20 micromolar/min 40 micromlar/min 70](https://cdn.numerade.com/ask_images/6da3d8d26a2d4ae29bd1f8304a3135e3.jpg)
SOLVED: For an enzyme following Michaelis-Menten kinetics, the following Vo vs. S data were obtained: [S] 1 micromolar 2 micromolar 4 micromolar 8 micromolar Vo 10 micomolar/min 20 micromolar/min 40 micromlar/min 70
![Convert nanomolar [nM] to micromolar [μM] • Molar Concentration Converter • Hydraulics — Fluids • Compact Calculator • Online Unit Converters Convert nanomolar [nM] to micromolar [μM] • Molar Concentration Converter • Hydraulics — Fluids • Compact Calculator • Online Unit Converters](https://www.translatorscafe.com/static/ucvt/img/Mole-Baloons.jpg)
Convert nanomolar [nM] to micromolar [μM] • Molar Concentration Converter • Hydraulics — Fluids • Compact Calculator • Online Unit Converters
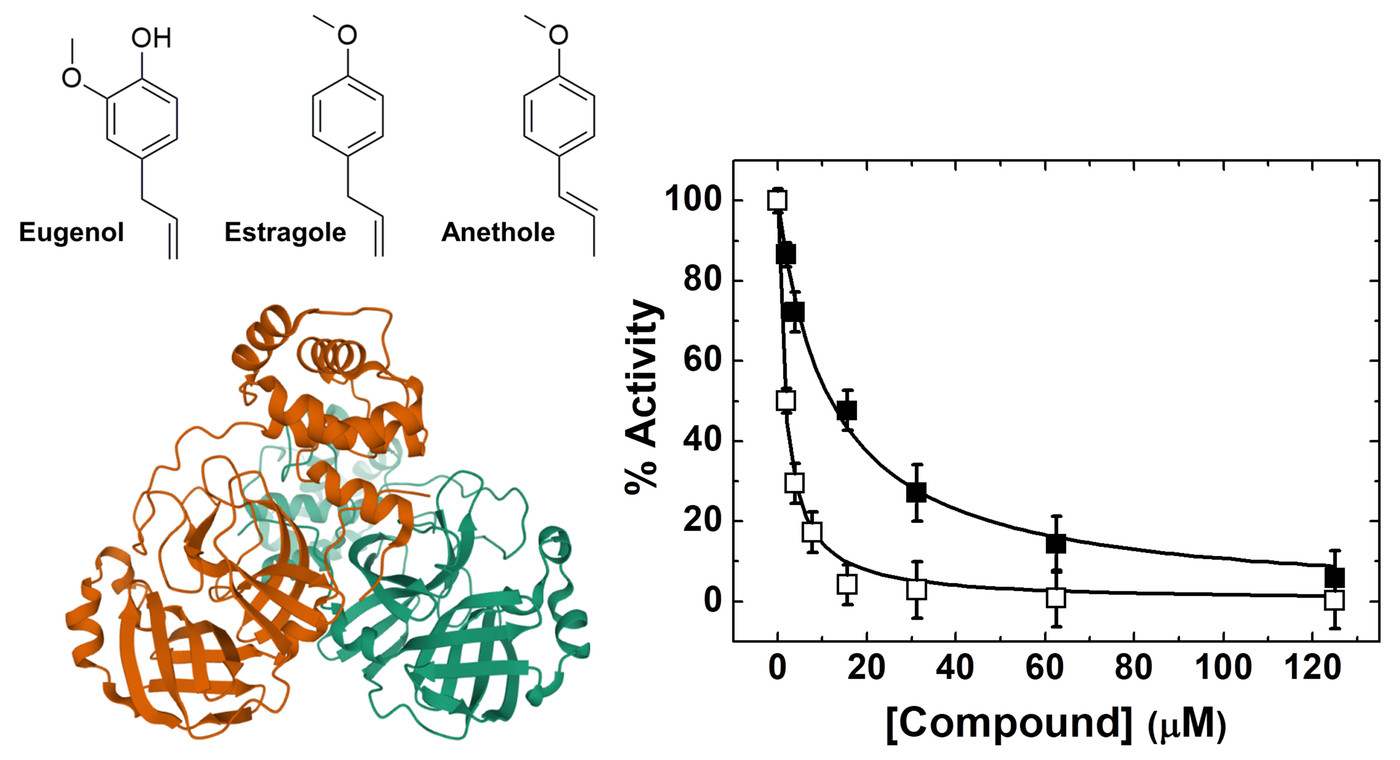
Pharmaceuticals | Free Full-Text | Sub-Micromolar Inhibition of SARS-CoV-2 3CLpro by Natural Compounds

CHARMM-GUI Free Energy Calculator for Practical Ligand Binding Free Energy Simulations with AMBER | Journal of Chemical Information and Modeling
PIC50: An open source tool for interconversion of PIC50 values and IC50 for efficient data representation and analysis

16 Molarity calculations ideas | converting metric units, measurement conversions, model question paper